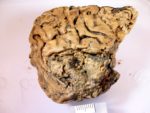
 In August of 2008, an archaeological survey in advance of new construction on the University of York’s Heslington campus discovered a human skull still containing brain remnants in a waterlogged pit. Surviving brain tissue is extremely rare, but it has been found before in archaeological contexts with extraordinary preservation of soft tissues — animals and people in the permafrost, bog bodies, desert mummies, crypt burials. These finds had other surviving soft tissues, however. The Heslington brain was unique as it is the only surviving soft tissue in otherwise fully skeletonized remains consisting of a skull, mandible and a couple of vertebrae. It is also the world’s oldest known surviving grey matter.
In August of 2008, an archaeological survey in advance of new construction on the University of York’s Heslington campus discovered a human skull still containing brain remnants in a waterlogged pit. Surviving brain tissue is extremely rare, but it has been found before in archaeological contexts with extraordinary preservation of soft tissues — animals and people in the permafrost, bog bodies, desert mummies, crypt burials. These finds had other surviving soft tissues, however. The Heslington brain was unique as it is the only surviving soft tissue in otherwise fully skeletonized remains consisting of a skull, mandible and a couple of vertebrae. It is also the world’s oldest known surviving grey matter.
 The location where the brain was found had a spring that had been source for wells from the Bronze Age through the middle Iron Age when the site was continuously occupied. In one of a dozen djacent pits apparently used for ceremonial offerings, archaeologists recovered a darkened cranium with articulated mandible face-down in moist sandy clay. The contents, first thought to be silt, were observed through the foramen magnum, inspected through an endoscope, X-rayed and a sample examined. Researchers confirmed that there was actual brain in that there skull.
The location where the brain was found had a spring that had been source for wells from the Bronze Age through the middle Iron Age when the site was continuously occupied. In one of a dozen djacent pits apparently used for ceremonial offerings, archaeologists recovered a darkened cranium with articulated mandible face-down in moist sandy clay. The contents, first thought to be silt, were observed through the foramen magnum, inspected through an endoscope, X-rayed and a sample examined. Researchers confirmed that there was actual brain in that there skull.
 The remains were radiocarbon dated to between 673-482 B.C. when the individual was struck hard on the head or neck and then decapitated, as evidenced by perimortem damage to the vertebrae and skull. The head was thrown in the pit and the brain had been naturally preserved in the waterlogged environment. It shrank over the millennia, but was still soft and glistening with clearly recognizable folds. Through cracks in the brain’s surface, the brain’s interior revealed itself to be a beige material with a tofu-like texture. Raman spectroscopic analysis found its chemical makeup to be largely decayed protein with a small amount of fat. Raman spectroscopy also identified the biochemical signatures of pigments produced by cyanobacteria. Researchers theorized that the cyanobacteria may have played a role in the unusual preservation of the brain tissue.
The remains were radiocarbon dated to between 673-482 B.C. when the individual was struck hard on the head or neck and then decapitated, as evidenced by perimortem damage to the vertebrae and skull. The head was thrown in the pit and the brain had been naturally preserved in the waterlogged environment. It shrank over the millennia, but was still soft and glistening with clearly recognizable folds. Through cracks in the brain’s surface, the brain’s interior revealed itself to be a beige material with a tofu-like texture. Raman spectroscopic analysis found its chemical makeup to be largely decayed protein with a small amount of fat. Raman spectroscopy also identified the biochemical signatures of pigments produced by cyanobacteria. Researchers theorized that the cyanobacteria may have played a role in the unusual preservation of the brain tissue.
 Now a new study of the brain’s proteins has revealed more information about its condition that may have long-term implications for medical research, particularly as regards brain diseases characterized by protein changes like dementia, Alzheimer’s, Huntington’s and Parkinson’s. Using electron scanning microscopes, University College London researchers were able to identify and unfold the brain’s preserved protein structures.
Now a new study of the brain’s proteins has revealed more information about its condition that may have long-term implications for medical research, particularly as regards brain diseases characterized by protein changes like dementia, Alzheimer’s, Huntington’s and Parkinson’s. Using electron scanning microscopes, University College London researchers were able to identify and unfold the brain’s preserved protein structures.
The researchers from University College London (UCL) show that of those substances which hold a human brain together, notably proteins, can fold themselves tightly into very stable structures, called aggregates.
Once unfolded – a process which [on the Heslington brain] took one year – these proteins regain many of the features typically encountered in a normal, living human brain. Scientists say the findings have implications for palaeoproteomics, biomarker research and diseases related to protein folding and aggregate formation.
Lead author Dr Axel Petzold, of the UCL Queen Square Institute of Neurology, had spent years researching two types of filaments in the brain – neurofilaments and glial fibrillary acidic protein (GFAP) – which act like scaffolds to hold brain matter together. He and his team found both of these were still present in the Heslington brain, suggesting they played a key role in keeping the brain matter together.
Typically, brains decompose quite quickly after death in a rapid process of autolysis – enzymes breaking up the tissue. The findings suggest that an acidic fluid may have got into the brain and prevented autolysis. Both filaments are typically found in greater concentrations in inner areas of the brain, but in the preserved Heslington brain there were more in the outer areas of the brain.
According to the researchers, this suggests the inhibition of autolysis would have started in the outer parts of the brain, potentially as an acidic fluid seeped into it.
The study has been published in the Journal of the Royal Society Interface.