 The body of a canid puppy preserved for 14,000 years in the Siberian permafrost has helped scientists make a crucial breakthrough in archaeological RNA analysis. In the right conditions, DNA can be extracted from archaeological remains thousands of years old, but the widely accepted hypothesis was that RNA degrades relatively quickly due to enzymatic action in plants and animals, especially in mammalian soft tissues. Recent studies have been able to reveal RNA genomes in archaeological material, almost all of the specimens from plant seed endosperm which is uniquely suited to long-term preservation. A new study sought to sequence RNA in historic and archaeological tissue specimens that, through rapid desiccation or freezing, had been exceptionally well-preserved.
The body of a canid puppy preserved for 14,000 years in the Siberian permafrost has helped scientists make a crucial breakthrough in archaeological RNA analysis. In the right conditions, DNA can be extracted from archaeological remains thousands of years old, but the widely accepted hypothesis was that RNA degrades relatively quickly due to enzymatic action in plants and animals, especially in mammalian soft tissues. Recent studies have been able to reveal RNA genomes in archaeological material, almost all of the specimens from plant seed endosperm which is uniquely suited to long-term preservation. A new study sought to sequence RNA in historic and archaeological tissue specimens that, through rapid desiccation or freezing, had been exceptionally well-preserved.
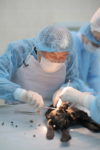
 The team took five samples from three canids: one sample each from two historical wolves (19th and 20th century) from Greenland, and tissue from the liver, cartilage and muscle of a “wolf” puppy discovered in Tumat, Siberia, in 2015. The pup was found in the thawing permafrost and many of its soft tissues had survived the 14,000 years in excellent condition. It was identified as a juvenile canid, but it’s not clear whether it was a wolf pup or a domesticated wolf-dog hybrid.
The team took five samples from three canids: one sample each from two historical wolves (19th and 20th century) from Greenland, and tissue from the liver, cartilage and muscle of a “wolf” puppy discovered in Tumat, Siberia, in 2015. The pup was found in the thawing permafrost and many of its soft tissues had survived the 14,000 years in excellent condition. It was identified as a juvenile canid, but it’s not clear whether it was a wolf pup or a domesticated wolf-dog hybrid.
 The DNA of the samples had already been sequenced, which gave researchers the means to verify the authenticity of any RNA results. Not surprisingly, there was much more RNA in the two historical samples, but the team was able to sequence the RNA from the Tumat canid’s liver tissue. This is the oldest directly sequenced RNA by at least 13,000 years.
The DNA of the samples had already been sequenced, which gave researchers the means to verify the authenticity of any RNA results. Not surprisingly, there was much more RNA in the two historical samples, but the team was able to sequence the RNA from the Tumat canid’s liver tissue. This is the oldest directly sequenced RNA by at least 13,000 years.
“Ancient DNA researchers have previously been reluctant to attempt to sequence ancient RNA because it is generally more unstable than DNA, and more prone to enzymatic degradation,” [University of Copenhagen’s Dr. Oliver] Smith said.
“However, following our recent successes in sequencing ancient RNA from plant material, we speculated that a well-preserved animal specimen, frozen in the permafrost, just might retain enough material to sequence.”
“To our delight, we found that not only did we find RNA from various tissues, but in some case the signal was so strong that we could distinguish between tissues in a way that makes biological sense.”
“Knowing that RNA acts as an intermediary between DNA and proteins, both of which are more stable, it might be tempting to ask, ‘so what?’ But we think the future of ancient RNA has great potential. For example, many of the most clinically relevant viruses around today have RNA genomes, and the RNA stage is often crucial to understanding the intricacies and complexities of gene regulation. This might have repercussions when discussing the environmental stresses and strains that drive evolution.”
The results of the study have been published in the open access journal PLoS Biology.